Yeast microfluidics
A case study on using Biocellion for designing microfludic chips for single cell analysis. In multiscale, agent based modeling, single cell analysis will be an important methodology for identifying cellular properties and the in silico ‘rules’ that lead to emergent behaviors of the multicellular system. Conversely, this experimental analysis can also benefit from computational modeling studies. In a collaboration between the Institute for Systems Biology (ISB) in Seattle WA USA and Institute of Genetics and Molecular and Cellular Biology (IGBMC) in Illkirch-Graffenstaden France scientists are using Biocellion to model the behavior of dividing yeast within microfluidic chips. The immediate goal is to help intelligently design new chips that improve the efficiency and speed of single cell analysis. Furthermore, development of general strategies that promote a symbiotic relationship between experimental and computational research will be critical in advancing our knowledge of complex biological systems.
One of the most exciting aspects of multiscale, agent based modeling is the study of emergent properties. These are properties that become manifest only at the level of the complete system, and are not obvious given the properties of the underlying parts. In cellular biology, we could imagine the underlying parts to be specific cells and the emergent properties to be population based phenotypes, such as the complex patterning of certain microbial colonies. Such patterns arise due to the behavior of the cells that make up the colony; however, the knowledge of a specific cell’s proliferation rate and/or other properties does not make the complex behavior of the colony immediately obvious. To study such a system in silico, we represent cells as agents and their behaviors guided by specific, logical rule sets. The agents can then be combined, in silico, to study various properties of the system. From this prospective it becomes obvious that proper formulation of the rules governing specific agents is paramount to the study of system phenotype and possible emergent properties. Currently, we are attempting to more systematically develop and validate specific rules governing key cellular processes. The rising interest in single cell analysis holds great promise in facilitating this goal. In such an analysis, the dynamics of individual cells are observed in various situations. Non-lethal approaches, such as fluorescence microscopy, even allow for the monitoring of intracellular molecular states. These experiments will allow us to better understand the cellular rules of proliferation, aging, death and response to various stimuli. Which will lead to the development of more accurate multiscale, agent based models.
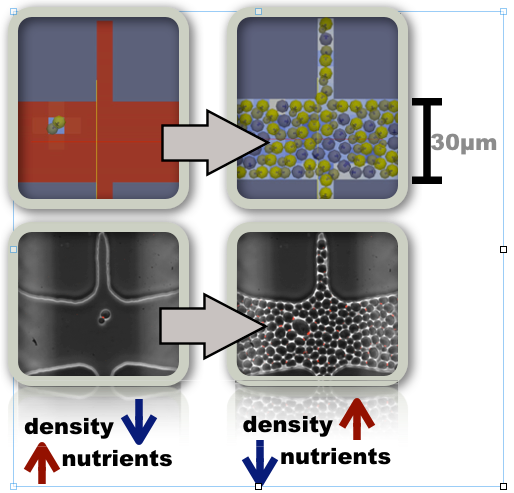
Importantly, the study of single cell analysis also stands to benefit from in silico computational modeling. Specifically, simulating the experimental system directly can help solve difficult engineering problems associated with experimental design and optimization. One example is the design of microfluidic chips to capture and study individual budding yeast cells. Researchers in Dr. Gilles Charvin’s Lab, at the Institute of Genetics and Molecular and Cellular Biology (IGBMC) in Strasbourg France, are currently implementing such strategies to better understand yeast aging dynamics, and have recently published their work in Cell Reports [1]. However, improving the throughput and developing larger and more complex systems has proven to be a difficult optimization problem involving various forces, filling dynamics and nutrient distribution. To solve this problem IGBMC is collaborating with ISB to develop high resolution multiscale, agent based models of the yeast-microfluidic system.
The current yeast-microfluidic prototype includes a validated model of yeast budding dynamics, which is coupled to the local environment by nutrient uptake. The agents interact with neighbors via shoving while mother bud pairs are held together via adhesion. Torque forces are also included to maintain the constant bud direction observed in actual cells. The distribution of nutrients in the environment is solved using PDEs with boundary conditions modeled after actual experimental conditions. Availability of nutrients is communicated to nearby agents, which in turn impacts growth rates. The geometry is modeled to reflect the actual geometry of the microfluidic chip, as well as multiple alternative designs and tunable parameters such as channel width. The spatial resolution is on the order of microns, and the time scales involved span hours, for the multiple divisions necessary to fill the chip, while maintaining proper accuracy for fast time scale events like nutrient diffusion. To achieve this level of detail, while maintaining reasonably fast simulation times, we have implemented this system in the Biocellion framework. Currently, simulations of more than 1000 cells can be simulated for 24 hours in less then 10 minutes on a single processor. This speed is achievable without sacrificing the high spatial and temporal resolution needed to model microfluidic systems. Investigation of the system to larger and more complex chips will benefit from Biocellions natural parallelization, as increased system size can actually enhance computational efficiency.
The first step is model validation, where simulation output is directly compared to multiple quantitative measures, such as: growth rates, mean squared translational and rotational displacement, diffusion patterns of fluorescent dyes and packing density estimates. The second step is optimization of larger more complex chip geometries which is done by searching over a large, high dimensional configuration space. Optimal candidates, identified by simulation, will then be fabricated and experimentally tested. The improved design will facilitate study of single cell behaviors, which will in turn, improve rules for more complex simulations such as yeast colony patterning.
1) Aging Yeast Cells Undergo a Sharp Entry into Senescence Unrelated to the Loss of Mitochondrial Membrane Potential; Fehrmann, Steffen; Paoletti, Camille; Goulev, Youlian; Ungureanu, Andrei; Aguilaniu, Hugo; Charvin, Gilles; Cell Reports doi:10.1016/j.celrep.2013.11.013 (volume 5 issue 6 pp.1589 - 1599)